Barriers to sequencing for everyday diagnostics
If it wasn’t clear before COVID-19, it is now: ubiquitous, worldwide testing for infectious diseases is crucial for public health and prosperity. Nucleic acid amplification tests (NAATs) are the gold standard for infectious disease diagnosis. Unfortunately, NAATs are a game of whack-a-mole: you have to know what disease you’re looking for in order to develop a test to diagnose it.
The promise of sequencing in the field
The solution that will inevitably displace conventional NAATs is metagenomic next generation sequencing (mNGS). Where each NAAT can detect only the specific primers the test is built around, mNGS reads the genome of the pathogen directly. Instead of a game of 20-questions (“Do you have flu? Do you have RSV? Do you have SARS?”), sequencing simply asks, “What do you have?” Over the last several years, researchers have been demonstrating the clinical utility of sequencing to identify any infectious disease.
If mNGS were ubiquitous, we could use the same tool for everyday diagnosis as we use to discover new diseases in the first place. But mNGS is not ubiquitous, because it’s time consuming and expensive.
That’s changing. Sequencing technology is cheaper and faster than ever, and it keeps getting smaller. Computational pipelines are being developed to process the complex data of mNGS in the cloud.
mNGS barriers: cost and complexity
In spite of these advances, routine mNGS testing remains limited by the cost and the complexity of pre-analytical preparation. Fieldable mNGS costs several thousand dollars, where NAATs cost less than $100. The mNGS sample and library prep workflows vary widely depending on the specific diagnostic workflow. For example, the pathogen sequence must be identified among the high background from host sequences. This means that the pathogenic material has to be preferentially lysed, purified or enriched. These methods that are typically complex and operator intensive.
The cost issue will fade in time. Within the decade, mNGS will cost less than $100 if costs continue to drop at the the historical rates. New technologies, such as Ontera’s NanoCounter, use silicon-based nanopores, which are manufactured using the same global semiconductor infrastructure behind computers and smartphones.
The mNGS workflow complexity issue, however, remains unsolved. The most practical solution for point-of-care sample and library preparation is a lab-in-a-suitcase, which has been used for Ebola detection in West Africa, and produced test results in under 24 hours. But the suitcase workflow is still operator-intensive, with significant hands-on time and opportunity for error and contamination, even with highly trained users.
Open questions
In short, sample and library prep are the major barriers to fieldable sequencing. These workflows are too complex and too prone to contamination or user error to hand off to relatively lesser trained operators and uncontrolled conditions in the field. Unless someone develops a magic-bullet chemistry that can get raw samples ready for sequencing in a “single pot” reaction, we’re stuck with automating the complex workflows used today. Historically, automation has been achieved with liquid handling systems, which are costly, complex, and expensive, not to mention impractical for fieldable analysis.
The path forward is to automate these workflows on a microfluidic cartridge. This is the way NAATs moved from liquid handling systems to point-of-care devices in the past decade. mNGS pre-analytical preparation will be harder to move to cartridges than NAATs were. Even today, developers of point-of-care NAAT platforms struggle to implement sample prep on-cartridges. Migrating NAATs to a cartridge is rife with challenges, and usually ends with compromised assay performance. The workflows for mNGS are more complex, the purity of the samples more important, and the diversity of applications are greater still.
It seems unlikely that the ideal pre-analytical prep will be a one-size-fits-all solution for mNGS. Instead, there is a need to develop modular solutions for reagent handling, purification, concentration, ligation, and amplification. Perhaps one day it will be as easy to design and assemble a customized sample-to-answer cartridge as it is to run a manual pipette-driven assay today.

400 Park Offices Dr.
Suite 301
RTP NC 27709
In The News
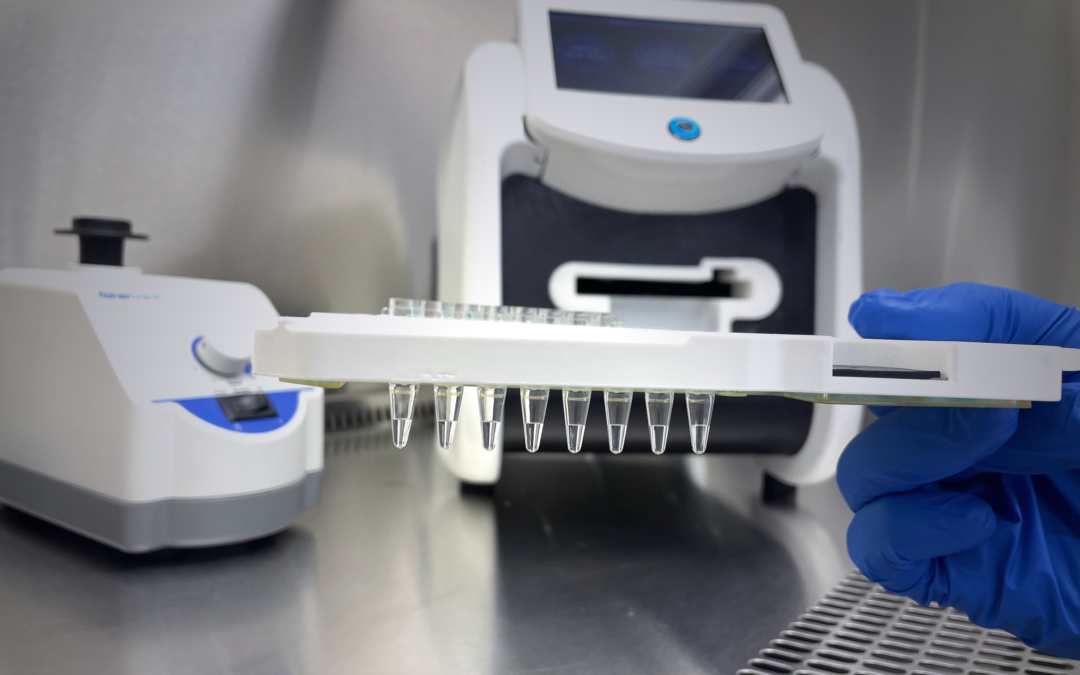
Product Update
Our newest cartridge and instrument is up and running. This marks a significant milestone: NAxtract now processes eight samples, where our prototype processed just
one. These new instruments are going through verification and validation now, with deliveries scheduled for early next year.
